|
The chromosomes of Gene Expression Programming are usually
composed of more than one gene of equal length. For each problem or
run, the number of genes, as well as the length of the head, are a
priori chosen. Each gene codes for a sub-ET and, in problems with
just one output, the sub-ETs interact with one another forming a
more complex multi-subunit ET; in problems with multiple outputs,
though, each sub-ET evolves its respective output.
Consider, for example, the following chromosome with length 39,
composed of three genes, each with length 13 (the tails are shown in
blue):
|
012345678901201234567890120123456789012 |
|
|
*Q-b/abbbaaba/aQb-bbbaabaa*Q-/b*abbbbaa |
(8) |
It has three open reading frames, and each ORF codes for a sub-ET
(Figure 6). We know already
that the start of each ORF coincides with the first element of the
gene and, for the sake of clarity, for each gene it is always
indicated by position zero; the end of each ORF, though, is only
evident upon construction of the corresponding sub-ET. As you can
see in Figure 6, the first
open reading frame ends at position 7; the second ORF ends at
position 3; and the last ORF ends at position 9. Thus, GEP
chromosomes contain several ORFs of different sizes, each ORF coding
for a structurally and functionally unique sub-ET. Depending on the
problem at hand, these sub-ETs may be selected individually
according to their respective outputs, or they may form a more
complex, multi-subunit expression tree and be selected as a whole.
In these multi-subunit structures, individual sub-ETs interact with
one another by a particular kind of posttranslational interaction or
linking. For instance, algebraic sub-ETs can be linked by addition
or subtraction whereas Boolean sub-ETs can be linked by OR or AND.
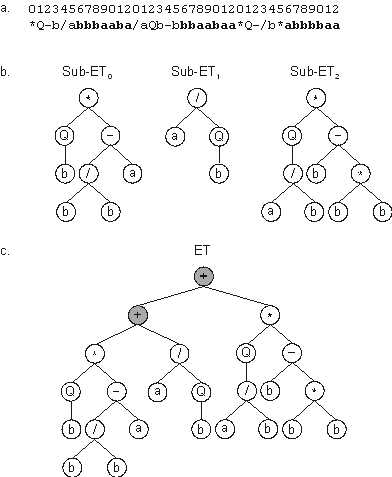
Figure 6. Expression of GEP genes as sub-ETs. a) A
three-genic chromosome with the tails shown in bold. Position zero marks the start of each gene.
b) The sub-ETs codified by each gene.
c) The result of posttranlational linking with addition. The linking functions are shown in gray.
The linking of three sub-ETs by addition is illustrated in
Figure 6, c. Note that the
final ET could be linearly encoded as the following K-expression:
|
012345678901234567890123 |
|
|
++**/Q-Q-aQ/b*b/ababbbbb |
(9) |
However, the use of multigenic chromosomes is more appropriate to
evolve good solutions to complex problems, for they permit the
modular construction of complex, hierarchical structures, where each
gene codes for a smaller and simpler building block. These building
blocks are physically separated from one another, and thus can
evolve independently. Not surprisingly, these multigenic systems are
much more efficient than unigenic ones (Ferreira 2001, 2002a).
And, of course, they also open up new grounds to solve problems of
multiple outputs, such as parameter optimization or classification
problems (Ferreira 2002a).
|
